You may have noticed a lack of activity on this blog. I moved to https://brendanhalpin.net/blog some time ago. Please check that location out instead!
Category Archives: Uncategorized
Web-scraping, survey data and Irish attitudes to immigration
This blog describes an ongoing research project which uses both survey data (from the European Social Survey) and web-scraping (of Google News data via SerpApi) to understand how Irish attitudes to immigration have changed over the last two decades.
The Republic of Ireland has undergone significant social transformations in the last few years, and one of the reasons for this transformation is immigration. Up until now, survey data and questionnaires from repositories like the Central Statistics Office (CSO), Eurobarometer, European Social Survey (ESS), and other public data sources have been used for different observations and analyses for studying societal attitudes. These sources provide first-order data that are gathered through efficient sampling strategies, and they update their data regularly, highlighting different topics and subtopics for that particular year.
Continue reading Web-scraping, survey data and Irish attitudes to immigrationThe role of indelcost: OM, LCS and Hamming
I will be blogging at https://brendanhalpin.net/blog/ from now on. Link to new post: https://brendanhalpin.net/blog/posts/omindel/
Parameterising TWED
I will be blogging at https://brendanhalpin.net/blog/ from now on. Link to new post: https://brendanhalpin.net/blog/posts/parameterisingtwed
Irish Universities’ Catchments: new data
Prompted by the arrival of another year’s “Feeder School” data in the Irish Times recently, I decided to update an analysis I have done in the past (Cullinan and Halpin, 2017), looking at the catchments of the several univerisities in Ireland. The picture I had from the previous analysis was that Galway, UL and UCD were most national in their recruitment patterns, TCD rather less so and UCC very markedly local.
Continue reading Irish Universities’ Catchments: new dataNetwork Sampling Blueskytes
I’m curious about Bluesky, and have accidentally found I can access part of its API. In particular, given a user’s handle, I can download their skytes, their posts. I got tipped off here: https://mastodon.social/@badlogic@mastodon.gamedev.place/111246801418649849
Continue reading Network Sampling BlueskytesWriting a Stata Command
In an idle moment this afternoon, I wrote a Stata command.
It was to create a light-weight implemention of the “percentogram” described at https://statmodeling.stat.columbia.edu/2023/04/13/the-percentogram-a-histogram-binned-by-percentages-of-the-cumulative-distribution-rather-than-using-fixed-bin-widths/, and I like the result, but it struck me that it is a good example of how practical and useful it can be to engage in Stata programming. Also, it’s a good example of how writing code in Stata (in a programmable command language) is very different from writing code in a stats-capable programming language like R, Python or Julia.
Continue reading Writing a Stata CommandTwitter activity after the ElMuskalypse
Premise
Did Twitter lose activity since the ElMuskalypse? Is my timeline quieter than it used to be when I was reading it? I mothballed my account around the end of November, after Elon Musk took over. How much activity have I been missing? To what extent have the people I followed also stepped back from Twitter?
How would you measure activity of your Twitter (ex-)timeline? Using the Twitter API, how would you assess whether the people you follow are more or less active? The simplest idea is to download the tweeting history of everyone you followed, and tot up their tweets by day. In principle that’s easy to do, if you have access to the Twitter API (and it still works). But it turns out it’s a bit more complicated than that, if you want to use this data to characterise how your timeline would have behaved in the interim.
Continue reading Twitter activity after the ElMuskalypsePremise
Did Twitter lose activity since the ElMuskalypse? Is my timeline quieter than it used to be when I was reading it? I mothballed my account around the end of November, after Elon Musk took over. How much activity have I been missing? To what extent have the people I followed also stepped back from Twitter?
How would you measure activity of your Twitter (ex-)timeline? Using the Twitter API, how would you assess whether the people you follow are more or less active? The simplest idea is to download the tweeting history of everyone you followed, and tot up their tweets by day. In principle that’s easy to do, if you have access to the Twitter API (and it still works). But it turns out it’s a bit more complicated than that, if you want to use this data to characterise how your timeline would have behaved in the interim.
Continue reading Twitter activity after the ElMuskalypsePinging Mastodon instances
Mastodon is less discoverable than Twitter, particularly because it doesn’t have full-text search, and because it has multiple instances. How do we know what’s going on, or which instances are particularly active?
There is a webpage of instance data somewhere. That is a good starting point.
But what about propagation of individual posts? How do they get seen, how quickly do other instances pick them up, which other instances? Here I summarise a quick experiment that exploits a feaure of mastodon software’s behaviour to map out some details of how a post is seen by other instances. Continue reading Pinging Mastodon instances
Viral Variants
Scary mutant COVID-19
Since the news of a potentially more transmissible strain of C19 broke in the UK, I’ve been thinking about the mechanics.
I was initially sceptical because it seemed to help politicians evade blame. Given the earlier fuss about a strain spread by holidaymakers returning from Spain in the summer, how much of the emergence of a new strain was down to network effects (common because it was, for example, present in a number of super-spreader events) versus inherent infectiousness?
So I simulated, with a workbench of 160000 agents in a 2D grid network, 1600 seed infections, and daily contact between 23 of the nearest 24 neighbours, and one random remote case. Base infectiousness, death and recovery rates are set to yield an R0 of about 3 and a SIR plot as in Figure 1.
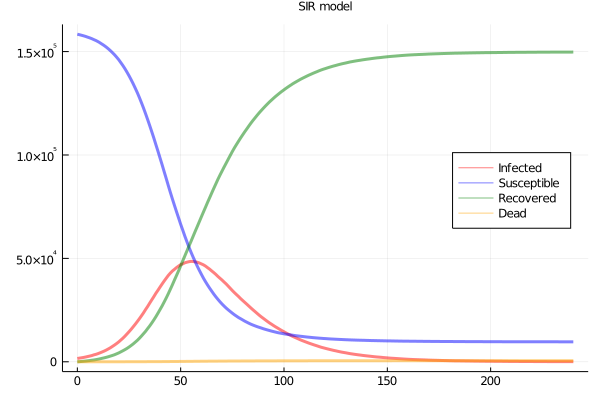
Figure 1: SIR plots